COVID-19 is not the only infectious disease New Zealand wants to eliminate, and genome sequencing is a crucial tool
Genome sequencing — the mapping of the genetic sequences of an organism — has helped track the spread of COVID-19 cases in Auckland, but it also plays an important role in the control of other infectious diseases in New Zealand.
One example is Mycoplasma bovis, a global cattle disease New Zealand also hopes to eliminate.
It was first detected on a South Island dairy farm in July 2017 and has subsequently been found on 250 properties across the country. It remains active on one farm.
M. bovis causes a range of diseases in both adult cattle and calves, including pneumonia, arthritis, mastitis, conjunctivitis and middle ear infections. The original source of the incursion remains unknown, but the Ministry for Primary Industries, together with industry partners, runs a programme to eliminate M. bovis from New Zealand.
Sequencing, epidemiology and evolutionary modelling all help determine how it spread between infected farms. Understanding “who infected whom” is crucial for contact tracing and control measures — and it applies whether a person is infected with COVID-19 or a cattle herd with M. bovis.
Read more: Eradicating cattle disease M. bovis in New Zealand may be costly, even impossible, but we must try
Eliminating human and animal pathogens
Several factors have contributed to the improvement in genome sequencing, including collaboration across New Zealand, an injection of new funding, and advances in modelling and visualisation.
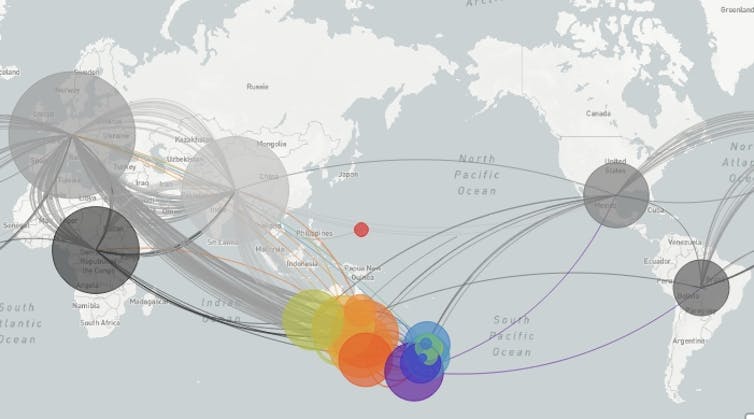 This map shows how COVID-19 genome sequences track transmission routes.
Nextstrain, supported by data from GISAID, CC BY-SA
This map shows how COVID-19 genome sequences track transmission routes.
Nextstrain, supported by data from GISAID, CC BY-SA
In the case of COVID-19, the turnaround speed from test sample to viral genome sequence has increased dramatically, and the information is invaluable for New Zealand’s ongoing effort to eliminate community transmission.
Many disciplines and skill sets are involved in analysing genome data and linking viral or bacterial sequences with epidemiological data on specific cases. New Zealand scientists work with international colleagues at the forefront of global projects to turn genome sequencing data into information for policy and action.
Read more: Why New Zealand needs to focus on genome sequencing to trace the source of its new COVID-19 outbreak
Apart from New Zealand’s effort to eliminate the cattle disease, other programs to control infectious diseases also benefit from genome sequencing. These include bacteria transmitted between animals and people, such as Salmonella, E. coli and Campylobacter.
Strategies to reduce food-borne Campylobacter infections in New Zealand are informed by “source attribution” models. These use bacterial sequences from human cases and animal hosts to determine the likely origin of human infection.
This information has helped the Ministry for Primary Industries and the food industry develop and implement policy to reduce the risk of food poisoning caused by Campylobacter.
From global to local control of infectious diseases
Genome sequencing, epidemiology and evolutionary modelling have combined to help understand transmission paths of infectious diseases at different scales. In New Zealand, these include the following:
In recent years, the Ministry of Health has funded the Institute of Environmental Science and Research to routinely sequence bacteria that cause disease in people. This has supported the control of outbreaks of Salmonella, E. coli, bacterial meningitis and antibiotic-resistant bacteria.
New research at the New Zealand Food Safety Science and Research Centre combines farm and factory-scale micro-mapping with genomics and modelling to control bacteria such as Listeria and Campylobacter. This helps our understanding of how microbes enter the food chain and how we can control them at the source.
Genome sequencing of new epidemics
We live in challenging times. We receive daily reminders of the consequences of our ability, or inability, to prevent and control emerging pathogens. Genome sequencing and modelling provide powerful opportunities to improve the management of infectious diseases globally.
It is exciting and encouraging to witness these advances in New Zealand. Recent investment in the country’s sequencing capability has led to an unprecedented acceleration in turnaround times. This makes the findings far more useful at the frontline of a rapidly evolving outbreak.
To reduce the impact of emerging diseases such as COVID-19, we need sustained investment in these new technologies. Human and animal infectious disease experts need to work together, along with experts in public health, microbiology, molecular biology, epidemiology and modelling.
We need to grow capability and maintain scientific networks to respond rapidly and effectively when the next disease crosses our borders.
Authors: Nigel French, Professor of Food Safety and Veterinary Public Health, Massey University





